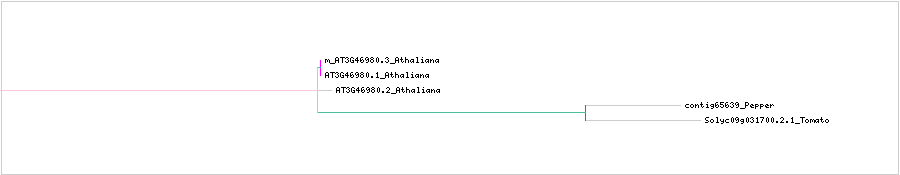
CLUSTAL W (1.83) multiple sequence alignment
m_AT3G46980.3_Athaliana MCYSLSIQSSIDFHNRNALKIHGDRAILTSNLPTLRRIPFLPERDRRRKL
AT3G46980.1_Athaliana MCYSLSIQSSIDFHNRNALKIHGDRAILTSNLPTLRRIPFLPERDRRRKL
AT3G46980.2_Athaliana MCYSLSIQSSIDFHNRNALKIHGDRAILTSNLPTLRRIPFLPERDRRRKL
contig65639_Pepper ------------MNSPASLRYLGPTSSSFRKPDSFGRRLVFKVKLPSNCK
Solyc09g031700.2.1_Tomato -MQIIFNNTTAQMNSPATLRYLS-TSSSFRKPNFFAKQLVFNDKLPSSCK
::. :*: . : : : : .: :
m_AT3G46980.3_Athaliana VLCTGRVVNSLKFTGNTSVDLCGIPRHRLRVSCSDARRTPEETAAELTAQ
AT3G46980.1_Athaliana VLCTGRVVNSLKFTGNTSVDLCGIPRHRLRVSCSDARRTPEETAAELTAQ
AT3G46980.2_Athaliana VLCTGRVVNSLKFTGNTSVDLCGIPRHRLRVSCSDARRTPEETAAELTAQ
contig65639_Pepper FSSNRNSF---AANSFFRREFSARRSYRIWGIEASSNDSVSVASGRIKTE
Solyc09g031700.2.1_Tomato FPPNHKSFKLITANTFFRTKIPVR-SNRIR---TSSNDSQSVQQ-----Q
. . . . . .: *: :.:. : . :
m_AT3G46980.3_Athaliana PNFSEFITSERVKVVAMLALALALCNADRVVMSVAIVPLSLSRGWSKSFS
AT3G46980.1_Athaliana PNFSEFITSERVKVVAMLALALALCNADRVVMSVAIVPLSLSRGWSKSFS
AT3G46980.2_Athaliana PNFSEFITSERVKVVAMLALALALCNADRVVMSVAIVPLSLSRGWSKSFS
contig65639_Pepper ESFVEFITSERVKVVAMLAVSLSLCNADRVVMSVAIVPLSLSHGWRQSFA
Solyc09g031700.2.1_Tomato GSFVEFITSERVKVVAMLALSLALCNADRVVMSVAIVPLSLSHGWRQSFA
.* ***************::*:*******************:** :**:
m_AT3G46980.3_Athaliana GIVQSSFLWGYLISPIAGGTLVDRYGGKVVMAWGVALWSLATFLTPWAAD
AT3G46980.1_Athaliana GIVQSSFLWGYLISPIAGGTLVDRYGGKVVMAWGVALWSLATFLTPWAAD
AT3G46980.2_Athaliana GIVQSSFLWGYLISPIAGGTLVDRYGGKVVMAWGVALWSLATFLTPWAAD
contig65639_Pepper GVVQSSFLWGYLISPIAGGTLVDYYGGKLVMAWGVALWSLATLLTPWAAE
Solyc09g031700.2.1_Tomato GVVQSSFLWGYLISPIAGGTLVDYYGGKLVMAWGVALWSMATLLTPWAAE
*:********************* ****:**********:**:******:
m_AT3G46980.3_Athaliana SSLWALLAARAMVGVAEGVALPCMNNMVARWFPPTERSRAVGIAMAGFQL
AT3G46980.1_Athaliana SSLWALLAARAMVGVAEGVALPCMNNMVARWFPPTERSRAVGIAMAGFQL
AT3G46980.2_Athaliana SSLWALLAARAMVGVAEGVALPCMNNMVARWFPPTERSRAVGIAMAGFQL
contig65639_Pepper ASLWALLAMRMLLGIAEGVALPCMNNMIARWFPPTERSRAVGLAMAGFQL
Solyc09g031700.2.1_Tomato VSLWALLAMRMLLGIAEGVALPCMNNMIARWFPPTERSRAVGLAMAGFQL
******* * ::*:************:**************:*******
m_AT3G46980.3_Athaliana GNVVGLMLSPILMSQGGIYGPFVIFGLSGFLWLLVWLSATSSAPDRHPQI
AT3G46980.1_Athaliana GNVVGLMLSPILMSQGGIYGPFVIFGLSGFLWLLVWLSATSSAPDRHPQI
AT3G46980.2_Athaliana GNVVGLMLSPILMSQGGIYGPFVIFGLSGFLWLLVWLSATSSAPDRHPQI
contig65639_Pepper GSAIGLTLSPILMSQGGLFGPFVIFGFSGFLWVLVWVSATSSAPECNWQI
Solyc09g031700.2.1_Tomato GSAIGLTFSPILMSQGGLFGPFVIFGLSGFLWVLVWVSATSSTPEHSRQI
*..:** :*********::*******:*****:***:*****:*: **
m_AT3G46980.3_Athaliana TKSELEYIKQKKQIS-TMENKRISTSGIPPFGRLLSKMPTWAVIVANSMH
AT3G46980.1_Athaliana TKSELEYIKQKKQIS-TMENKRISTSGIPPFGRLLSKMPTWAVIVANSMH
AT3G46980.2_Athaliana TKSELEYIKQKKQIS-TMENKRISTSGIPPFGRLLSKMPTWAVIVANSMH
contig65639_Pepper SIHELMYVQNKGQIHTIAEGKSKTSKGIPPFRCLLSKLPTWSIIVANAMH
Solyc09g031700.2.1_Tomato SIHELRYIQNKGQIHNIVEGKSKSSKGIPPFRRLLSKLPTWSIIVANAMH
: ** *:::* ** *.* ::.***** ****:***::****:**
m_AT3G46980.3_Athaliana SWGFFVILSWMPIYFNSVYHVNLKQAAWFSAVPWSMMAFTGYIAGFWSDL
AT3G46980.1_Athaliana SWGFFVILSWMPIYFNSVYHVNLKQAAWFSAVPWSMMAFTGYIAGFWSDL
AT3G46980.2_Athaliana SWGFFVILSWMPIYFNSVYHVNLKQAAWFSAVPWSMMAFTGYIAGFWSDL
contig65639_Pepper SWGFFVILSWMPIYFKTIYHVDLRQAAWFSAVPWCMMALTGYFAGVLSDM
Solyc09g031700.2.1_Tomato SWGFFVILSWMPIYFKTIYHVDLRQAAWFSAVPWSMMALTGYFAGFLSDM
***************:::***:*:**********.***:***:**. **:
m_AT3G46980.3_Athaliana LIRRGTSITLTRKIMQSIGFIGPGIALIGLTTAKQPLVASAWLSLAVGLK
AT3G46980.1_Athaliana LIRRGTSITLTRKIMQSIGFIGPGIALIGLTTAKQPLVASAWLSLAVGLK
AT3G46980.2_Athaliana LIRRGTSITLTRKIMQSIGFIGPGIALIGLTTAKQPLVASAWLSLAVGLK
contig65639_Pepper MIQRGISVTLTRKVMQSIGFFGPGFALIGLTMAPTPSIASAWLTLAVGLK
Solyc09g031700.2.1_Tomato MIQRGISVTLTRKVMQSVGFFGPGFSLIGLTTAPSPSIASAWLTLAVGLK
:*:** *:*****:***:**:***::***** * * :*****:******
m_AT3G46980.3_Athaliana SFSHLGFLINLQEIAPEYSGVLHGMCLTAGTLAAIVGTVGAGFFVELLGS
AT3G46980.1_Athaliana SFSHLGFLINLQEIAPEYSGVLHGMCLTAGTLAAIVGTVGAGFFVELLGS
AT3G46980.2_Athaliana SFSHLGFLINLQVPK----RVIHT--------------------------
contig65639_Pepper AFSHCGFLVNLQEIAPQYSGVLHGLSNTAGTFAAIIGTVGAGFFVELVGS
Solyc09g031700.2.1_Tomato AFSHCGFLVNLQEIAPQYSGVLHGYKAAR--------PVKSSWHHCQC--
:*** ***:*** *:*
m_AT3G46980.3_Athaliana FQGFILLTAILYLLSALFYNIYATGERVDFDTTAA
AT3G46980.1_Athaliana FQGFILLTAILYLLSALFYNIYATGERVDFDTTA-
AT3G46980.2_Athaliana -----------------------------------
contig65639_Pepper FKGFLLLTASLYFSAAVFWDVFSTGERIKFDDST-
Solyc09g031700.2.1_Tomato --CCDLLTP---FFRLSFWYIFLLG----------
|