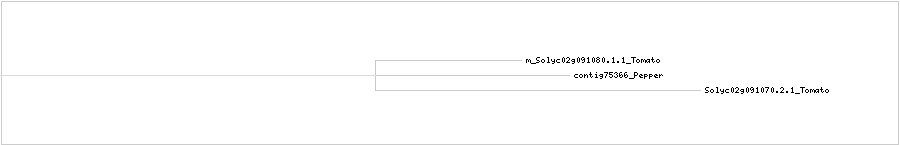
CLUSTAL W (1.83) multiple sequence alignment
m_Solyc02g091080.1.1_Tomato MKGTIDVTPDTKKLLVGIYSSIEITTSAAYSCRAESMEEQLLESEENRKW
contig75366_Pepper --------------------------------------------------
Solyc02g091070.2.1_Tomato -----------------------------------MEEELPQSLKLEKKW
m_Solyc02g091080.1.1_Tomato IITTEWETYCQELKKLSYIAAPMVAVSVLQYLLQVVSMMMVGHLDQISLS
contig75366_Pepper --------------------------------------------------
Solyc02g091070.2.1_Tomato QIS--WDALSQELKKTNRIVAPMVAVTVFQYLLQVVSIIMVGHLGQLALS
m_Solyc02g091080.1.1_Tomato SVSVATSITNVTGFSLLSGLVGGLETLSGQAYGANQWQKVGTYTHSAVIS
contig75366_Pepper --------------------------------------------------
Solyc02g091070.2.1_Tomato SVAIATSLTNVTGFSLLSGLVGGLETLCGQAYGAKQYHKLSTYTYTAIIS
m_Solyc02g091080.1.1_Tomato LLIVCIPISVVWLFMDKLLILMGQDPVISVEAYKFSLWLIPALFGSAILR
contig75366_Pepper --------------------------------------------------
Solyc02g091070.2.1_Tomato LFLVCLPICVLWFFMDKLLILIGQDHEISIEARKYALWAIPALFGGAISK
m_Solyc02g091080.1.1_Tomato PLLRYLLIQSLIFPMVVSAFLALCLHISLSWLFIFHLGLGKSGAAIAFSF
contig75366_Pepper --------------------------------------------------
Solyc02g091070.2.1_Tomato PLVRYLQTQSLILPMLLSSLAVLCLHLPISWGFIFKLELGNIGAAIAFSV
m_Solyc02g091080.1.1_Tomato SIWTFVASLVVYISRSSSCERTRTPLNRDAFLAIGQFFRYATPSALMVCL
contig75366_Pepper --------------------------------------------------
Solyc02g091070.2.1_Tomato STWLYVLFLALHVSFSTSCEKTRTPFSMDVFLGINEFFRLAVPSAVMVCL
m_Solyc02g091080.1.1_Tomato KWWSLEVLTLLSGLLPNPKLETSVLSICLTISALHFALPYGLGTGVSTRV
contig75366_Pepper --------------------------------------------------
Solyc02g091070.2.1_Tomato KWWSLEVLTLLSGLLPNPTLEASVLSICLTISTLHFTIPYGFGAAASTRV
m_Solyc02g091080.1.1_Tomato SNELGSGNPQKALVAVRVAMFVAVSETVLASIVIFLCRGVLGKAYSNDEQ
contig75366_Pepper -------------------MILAVAETIVASTVTFLCREVLGKVYSNDQQ
Solyc02g091070.2.1_Tomato SNELGAGNPKRARMAVQVVMFLAVIETVVLSTSMFGSRHILGRGFSNEKQ
*::** **:: * * .* :**: :**::*
m_Solyc02g091080.1.1_Tomato VVDYVARITPFLCLTIITDSFQAVICGVARGSGWQIIGAYVNLGAFYLVG
contig75366_Pepper VVDYVAAITPFLCLSIITDSLQGVISGIARGSGWQILGAYVNLGAFYLVG
Solyc02g091070.2.1_Tomato VVEYIASMTPLLCLSIITDSLQAVISGIARGSGWQHIGAYINLGAFYLVA
**:*:* :**:***:*****:*.**.*:******* :***:********.
m_Solyc02g091080.1.1_Tomato IPVAAVMCFLVNLKSKGLWIGLLAGSTIQATSLSLILVFTDWQKQAINAK
contig75366_Pepper IPVAAVLCFYVNLKAKGLWIGLLVGTAIQATSLSFILGFTDWQKQAIKAK
Solyc02g091070.2.1_Tomato IPLAVVLGFTLHMKAKGLWIGIVVGSTIQSIILSIVTCFTDWDKQAKKAR
**:*.*: * :::*:******::.*::**: **:: ****:*** :*:
m_Solyc02g091080.1.1_Tomato KRISERR-----------
contig75366_Pepper QRLSEKRSSVENDLEGDE
Solyc02g091070.2.1_Tomato ERILEGTS----------
:*: *
|