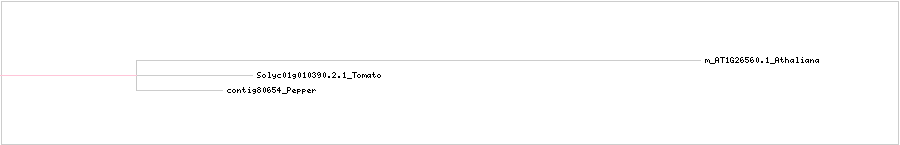
CLUSTAL W (1.83) multiple sequence alignment
Solyc01g010390.2.1_Tomato MLKKRGIDVWALIFLIFVISFINTNS--AQNFSRNNFPRDFVFGTASSAY
contig80654_Pepper MLKKRGIAIWSIFFILFVS--FDTNS--AQNISRNNFPKGFVFGTASSAY
m_AT1G26560.1_Athaliana MAHRRLIMTMTKMMMMVTMMMMMDKTCICADISRGSFPKGFVFGTASSAF
* ::* * : ::::.. : :: . ::**..**:.*********:
Solyc01g010390.2.1_Tomato QFEGAVKEDGRGQTIWDKFSHSFGKVIDFSNGDVAVDQYHRYLEDIQLMK
contig80654_Pepper QFEGAVKEDGRGQTIWDKFSHSFGKVIDFSNGDVAVDHYHRYPEDIQLMK
m_AT1G26560.1_Athaliana QHEGAVKAEGRGPTIWDTFSHTFGKITDFSNADVAVDQYHRYEEDVQLMK
*.***** :*** ****.***:***: ****.*****:**** **:****
Solyc01g010390.2.1_Tomato DMGMDAYRFSIAWARIFPNGTGEINQAGVDHYNKFINALLANGIKPYVTL
contig80654_Pepper DMGMDAYRFSIAWARIFPKGAGEINQAGVVHYNKLINALLANGIQPYVTL
m_AT1G26560.1_Athaliana NMGMDAYRFSISWTRIFPNGVGHINEAGIDHYNKLINALLAKGIEPYVTL
:**********:*:****:*.*.**:**: ****:******:**:*****
Solyc01g010390.2.1_Tomato YHWDLPQALEDKYTGWLSPQIIEDFAIYAETCFKKFGDRVKNWITINEPH
contig80654_Pepper YHWDLPQALQDKYNGWLSPQIIKDFAMYAETCFEKFGDRVKNWITINEPH
m_AT1G26560.1_Athaliana YHWDLPQALHDRYLGWLNPQIINDFAAYAEVCFQRFGDRVKHWITFNEPH
*********.*:* ***.****:*** ***.**::******:***:****
Solyc01g010390.2.1_Tomato TVAVQGFDVGLQAPGRCSILLRALCRAGNSATEPYIVTHNLLLAHATAVD
contig80654_Pepper TVAIQGFDVGLQAPGRCSILLRAFCRAGNSATEPYIVTHNLLLAHATVVD
m_AT1G26560.1_Athaliana TFAIQGYDVGLQAPGRCTILFKLTCREGNSSTEPYIVGHNVILTHATVSD
*.*:**:**********:**:: ** ***:****** **::*:***. *
Solyc01g010390.2.1_Tomato IYKKKYKPTQHGSIGISLDSFWYEPLTNSKDDIEATQRAIEFNLDWYLEP
contig80654_Pepper IYKKKYKPMQHGSIGISLDTFWYEPLTNSKDDIEATKRAIEFNLDWFLEP
m_AT1G26560.1_Athaliana IYRKKYKAKQGGSLGIAFDVMWFEPESNKTEDIEAAQRAQDFQLGWFLDP
**:****. * **:**::* :*:** :*..:****::** :*:*.*:*:*
Solyc01g010390.2.1_Tomato VILGRYPSSMVERVGSRLPKFSPTESALVKGSYDFIGINHYTTWYASKNK
contig80654_Pepper VMLGRYPSSMVDRVGSRLPKFSPAESALVKGSYDFIGINHYTTWYASKNK
m_AT1G26560.1_Athaliana LMFGDYPSSMRSRVGSRLPVFTGSQSSLVKGSLDFVGINHYTTYYARNNA
:::* ***** .******* *: ::*:***** **:*******:** :*
Solyc01g010390.2.1_Tomato TNIIGALLNDSVADSGAITLPFKGIKPIADRASSIWLYIVPSGIRSLLNY
contig80654_Pepper TNIIGVLLNDSVADSGAITLPFKGLKPIADRASSIWLYIVPQGIRSLMNY
m_AT1G26560.1_Athaliana TNLIGTLLHDAVSDSGTVTLPFKGLSTIGDRASSIWLYIVPRGMRSLMNY
**:**.**:*:*:***::******:..*.************ *:***:**
Solyc01g010390.2.1_Tomato IKQKYENPLIIITENGMDDANSIFTSREHTLKDTKRINYHNDYLTNLLDA
contig80654_Pepper IKQKYGNPLIIITENGMDDSNNVFTSRQDTLKDTKRISYHKDYLTNLLAA
m_AT1G26560.1_Athaliana IKHRYGNPPVFITENGMDDPNSILISRKDALKDAKRIKYHHDYLSSLQAS
**::* ** ::********.*.:: **:.:***:***.**:***:.* :
Solyc01g010390.2.1_Tomato IKEDGCNVKGYFVWSLMDNWEWAAGFSSRFGLYYVDYKDNLKRYPKDSVN
contig80654_Pepper IKEDGCNVKGYFVWSLMDNWEWAAGFSSRFGLYYVDYKDNLRRYPKDSVN
m_AT1G26560.1_Athaliana IKEDGCNVKGYFVWSLLDNWEWAAGYSSRFGLYFVDYRDNLKRYPKDSVH
****************:********:*******:***:***:*******:
Solyc01g010390.2.1_Tomato WFKNFLGSA-
contig80654_Pepper WFKNFLGSA-
m_AT1G26560.1_Athaliana WFTSFLNSTS
**..**.*:
|