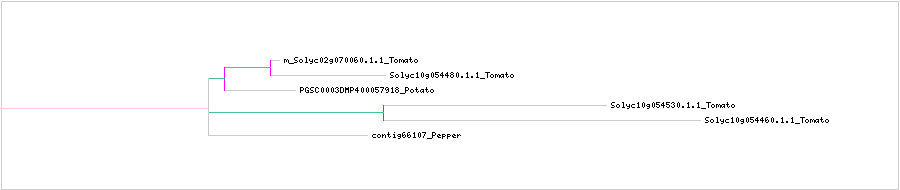
CLUSTAL W (1.83) multiple sequence alignment
m_Solyc02g070060.1.1_Tomato MVYCIVLLVFFLLTITPISYGNLQDFATCMSLHTTSTSSSVVHSQQSSSY
Solyc10g054480.1.1_Tomato -----------------------------MSFHTTSTSSSVVHSQQSSSY
PGSC0003DMP400057918_Potato MVYSIVLPVFFLLTITPISYENLQDFATCMSFHTTSTSSSVVHSQQSSSY
contig66107_Pepper -----------------------------MALHSTSTYSSVVHSQESPSY
Solyc10g054530.1.1_Tomato --------------------------------------------------
Solyc10g054460.1.1_Tomato -----------------------------MSLLWKMRFP--LRQVKCIMR
m_Solyc02g070060.1.1_Tomato TYLLQESQQNPRWLNSTSLLKPSFIVTPKTQNETQGAILCAKKHGLQVRV
Solyc10g054480.1.1_Tomato TYLLQESQQNPRWLNSTSLLKPSFIVTPKTQNEIQGAILCAKKDGLQIRV
PGSC0003DMP400057918_Potato TYLLQESQQNPRWLNSTSLLKPSFIVTPKTQNEIQGVILCAKKHGLQVRV
contig66107_Pepper AYLLQESQQNPRWLNSTSLLKPSFIVTPKTQNEIQGAILCAKKHSLQVRV
Solyc10g054530.1.1_Tomato --------------------------------------------------
Solyc10g054460.1.1_Tomato KYILKLKMEN--------LLRDSMVN-----------IICVN--------
m_Solyc02g070060.1.1_Tomato MSGGHDYEGLSYLCKKPFIILDLVDYRSINIDIENETAWIQSGATIGEVY
Solyc10g054480.1.1_Tomato MSGGHDYEGLSLLCKKPFIILDLVDYRSTNIDIENETAWIHSGATIGEVY
PGSC0003DMP400057918_Potato MSGGHDYEGLSFLCKNPFIILDLVDYRSINIDIENETAWIQSGATIGEVY
contig66107_Pepper MSGGHDYEGLSFLCKTPFIILDLIDYRSININLEDETAWIQSGATIGEVY
Solyc10g054530.1.1_Tomato --MGQEWKMYLFIY-----------------------------------W
Solyc10g054460.1.1_Tomato ------FHFLLLLTS-----------------------------------
:. :
m_Solyc02g070060.1.1_Tomato YNIAKKSNILGFPAGLCPSVGIGGHFSGGGIGTMMRKYGLAADNIIDANL
Solyc10g054480.1.1_Tomato YNIAKKSNILGFPAGLCPSVGVGGHLSGGGIGTMMRKYGMAADNIIDANL
PGSC0003DMP400057918_Potato YNIAKKSNILGFPAGLCPSVGVGGHFSGGGIGTMMRKYGLASDNIIDASF
contig66107_Pepper YNIAKKSDILGFPAGLCPSVGVGGHFSGGGIGTMMRKYGLSADNIIDANL
Solyc10g054530.1.1_Tomato FYILLK---------ICPSVGVGGHFSGGGIGTMMRKYDLAADNIIDANL
Solyc10g054460.1.1_Tomato --LLK----------ICPSVGVGGHFSGGGIGTMMRKYGLAADNIIDANL
: :*****:***:************.:::******.:
m_Solyc02g070060.1.1_Tomato VDAKGTILNRKTMGEDVFWAIRGGGGASFGVISAWKVKLVRVPSLVTVFT
Solyc10g054480.1.1_Tomato VDANGTILNRKTTGEDVFWAIRGGGGASFVVISAWKVRLVRVPSLVTVFT
PGSC0003DMP400057918_Potato VDANGRILNRKTMGEDVFWAIRGGGGASFGVISAWKVKLVRVPSLVTVFT
contig66107_Pepper VDANGRILNRETMGEDVFWAIRGGGGASFGVISAWKVRLVRVPPLVTVFT
Solyc10g054530.1.1_Tomato VDANGTILNRKTMGEDVFWAIRGGGGASFGVISAWKVRLVRVPSLVTVFT
Solyc10g054460.1.1_Tomato VDPNGTILNRKTMEEDVFWAIRGGGGASFGVISAWKVRLVRVSSLVTVFK
**.:* ****:* *************** *******:****..*****.
m_Solyc02g070060.1.1_Tomato IHKRLDQEGVELVHNWQYIASKLPEGLFIRVLIQQIDRIDGQGNVKLPEV
Solyc10g054480.1.1_Tomato IHKRLDQEGVELVHNWRYIASKLPEGLFIRVLIQQIDGIGKQGNVKLPEV
PGSC0003DMP400057918_Potato IHKRLDQEGVKLVHNWQYIASKLPKGLFIRVLIQQIDGVDSQGNVNQAEV
contig66107_Pepper IHKMLDQEGVKLVHKWQYIASKLPENLFIRVLIQQIDGVDSHGNVK---V
Solyc10g054530.1.1_Tomato IHKRLDQEGVELVHNWQYIANKLPEGLFIRVLIQQIDGIRSQGNVKLSEV
Solyc10g054460.1.1_Tomato IHKRLDQEGVELVHNWQYIASKLPEGLFIRVLIQQIDGIGRQGNVKLLEV
*** ******:***:*:***.***:.*********** : :***: *
m_Solyc02g070060.1.1_Tomato LFNSLFLGLKSDLISLMNANFPELGLKMEDCTEMSWIQSVLYFIGYKKGE
Solyc10g054480.1.1_Tomato LFNSLFLGLKSDLISLMNANFPELGLKMEDCTG-----------------
PGSC0003DMP400057918_Potato LFNSLFLGLKTDLISVMNTNFPELALKTEDCTEMSWIKSVLYLTGYQKGE
contig66107_Pepper LFNSLFLGLKTDVFSIMNTNFPELGLKMEDCAEMSWIRSVLYFTGYQKGE
Solyc10g054530.1.1_Tomato LFNSLFFGLKFDLISLMNANFPELGLKMEDCTEMSWIKSVLYFTGYQKGE
Solyc10g054460.1.1_Tomato LFNSLFLGLKSDLISLMNANFPELGLKMEDCTEMS---------------
******:*** *::*:**:*****.** ***:
m_Solyc02g070060.1.1_Tomato PLEVLLDRKTQYKSNFKAKSDFVVEPMPESVFQGISERFLHKKLVFMIMD
Solyc10g054480.1.1_Tomato --------------------------------------------------
PGSC0003DMP400057918_Potato PLEVLLDRKTQYKSNFKAKSDFVVEPMPESVFQGISERFLHQKLVFMIMD
contig66107_Pepper PLEILLDRTTQYKSNLKAKSDFVIEPMPESVFQGISERFSQQKMAFMIMD
Solyc10g054530.1.1_Tomato PLEVLLDRKAQYKSNFKVNSDFVVESMLESVFQGSQRGFFGKKKFS----
Solyc10g054460.1.1_Tomato --------------------------------------------------
m_Solyc02g070060.1.1_Tomato PLGGKMDEIEEYEIPFPHRKGNIYNIQYIVKWDSNEGSKQYLYWIQNLYK
Solyc10g054480.1.1_Tomato --------------------------------------------------
PGSC0003DMP400057918_Potato PLGGKMDEIDEYEIPFPHRKGNIYNIQYIVKWDSNEGSNEHLYWIQNLYK
contig66107_Pepper PLGGRMDEILESEIPFPHRKGNIYNIQYIVKWGSNEGSKLHLDWIQSLYK
Solyc10g054530.1.1_Tomato --------------------------------------------------
Solyc10g054460.1.1_Tomato --------------------------------------------------
m_Solyc02g070060.1.1_Tomato YMEPYVSNSPRASYLNYRDLDLGINQQGNDSSYGQAIMTWGTKYFKGNFQ
Solyc10g054480.1.1_Tomato --------------------------------------------------
PGSC0003DMP400057918_Potato YMEPYVSNSPRAAYISYRDLDLGINQQGNYSSYRQAIMTWGTKYFKGNFQ
contig66107_Pepper YMEPYVSNSPRSAYLNYRDLDLGINQQGNYLSYRQAIT-WGTKYFKGNFQ
Solyc10g054530.1.1_Tomato --------------------------------------------------
Solyc10g054460.1.1_Tomato --------------------------------------------------
m_Solyc02g070060.1.1_Tomato RLAKAKHQIDPNNFFTNEQSIPPLCC
Solyc10g054480.1.1_Tomato --------------------------
PGSC0003DMP400057918_Potato RLAKAKHQIDPNNFFTNEQSIPPLCC
contig66107_Pepper RLERAKHQIDPSNFFTNEQSIPPLSY
Solyc10g054530.1.1_Tomato --------------------------
Solyc10g054460.1.1_Tomato --------------------------
|